Verla Andrew Wirnkor1, Enyoh Christian Ebere1* and Verla Evelyn Ngozi2
1 Group Research in Analytical Chemistry, Environment and Climate change (GRACE&CC), Department of Chemistry, Faculty of Science, Imo State University, Owerri, Imo State Nigeria
2Department of Environmental Technology, School of Environmental Technology Federal University of Technology, Owerri, Imo State Nigeria.
*Email: This email address is being protected from spambots. You need JavaScript enabled to view it., +2347063715081
Abstract:
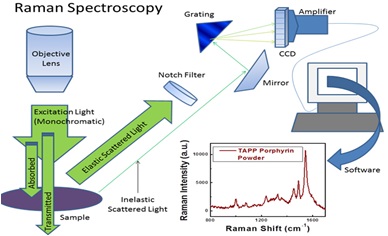
Microplastics are ubiquitous tiny plastic particles (< 5 mm) nonbiodegradable and have large surface area in the environment or the body of living things due to anthropogenic activities or fragmentation of plastic debris. Though they have been found in sea food and in human body, their health implications are still speculative. A major reason for death of information on this topical issue is the lack of standard methods for detecting and quantifying microplastics as a result of their presence in more complex environmental matrices. In the present review some methodologies for analyzing microplastics reported in the period 2000 to 2018 have been documented with the aim of assessing which methods is most suitable and in what matrix. The following methods have been studied: CHN analyzers, pyrolysis-gas chromatography/mass spectroscopy (PyrGC/MS), optical microscopy, fourier transform infrared microspectroscopy (Micro-FTIR), raman microspectroscopy (RMS) and scanning electron microscopy with energy dispersive x-ray spectroscopy (SEM-EDS). Studies have been conducted with often a combination of two methods; one separating and the other quantifying which can be problematic moreso in living tissue where there is no harm reported as at the time of this study. However, microplastics have become a cause for concern and advance studies are required to unravel the potential risk of their presence in our food and environment.
Keywords:
Anthropogenic, Fragmentation, Health implications, Hyphenated methods, Matrices, Seafood.

























![atomic absorption spectrophotometric determination of trace amount of copper in water samples after preconcentration with [n-[(s)-3-mercapto-2-methylpropiony]-l-proline] on a naphthalene atomic absorption spectrophotometric determination of trace amount of copper in water samples after preconcentration with [n-[(s)-3-mercapto-2-methylpropiony]-l-proline] on a naphthalene](/cache/mod_news_show_sp2/nssp2_thumbs/183/ATOMIC-ABSORPTION-SPECTROPHOTOMETRIC-DETERMINATION-OF-TRACE-AMOUNT-OF-COPPER-IN-WATER-SAMPLES-AFTER-PRECONCENTRATION-WITH-N-S-3-MERCAPTO-2--METHYLPROPIONYL-L-PROLINE-ON-A-NAPHTHALENE_170x230.jpg)